Flex Funds Project: MetaProt-KG
Metaproteomics-centric microbiome knowledge graph as tool for omics data integration and analysis (MetaProt-KG)
Project Start and End Date
2025-01-01 - 2025-12-31
Short project summary
Microbiomes are important for the environment, human health, biotechnology, and agriculture. Researching microbiomes involves multiple omics fields, particularly metagenomic, metaproteomic, and metabolomic. Crosslinking these domains is complex and time-consuming, requiring a systematic approach to streamline research and enhance data visibility.
Knowledge graphs (KG)s excel at representing and visualizing complex relationships between
entities and mapping intricate networks of genes, proteins, metabolites, and other biomolecules and metadata across life science domains. This holistic view is crucial for understanding mechanisms, regulatory networks, metabolic pathways, and identifying unknown proteins. KGs enable intuitive data exploration and hypothesis generation. Advanced computational techniques, such as machine learning and network analysis, leverage KGs to uncover patterns, predict functional roles, and identify key nodes in biological networks. LLMs and KGs synergize effectively, with LLMs using KGs as context to reduce hallucinations and enrich KGs by extracting and integrating the latest findings from vast amounts of unstructured biomedical literature. KGs also facilitate data sharing and collaboration across the scientific community, ensuring reproducibility and data quality. Their standardized structure maintains data provenance and quality, essential for robust research.
This project aims to create MetaProt-KG, a microbiome knowledge graph starting with metaproteomics, our area of expertise. The knowledge graph will be upgradable for all
microbiome research domains. The main objectives are:
- Creating an overview of all existing databases useful for metaproteomics
- Developing a metaproteomic knowledge graph from these databases deployed as a web tool on the de.NBI cloud.
- Creating a collection of queries and algorithms in a user-friendly way to analyse your data with KGs.
Finally, to maximize the impact of our proposed objectives, the knowledge graph will be
presented at conferences and workshops. By reaching a broader community, the long-term goal is to create a community knowledge graph encompassing more omics fields, more datasets, and standardized data formats. Furthermore, based on a general KG, individuals and institutions can create their instances or work on a combined community instance.
Graphical abstract
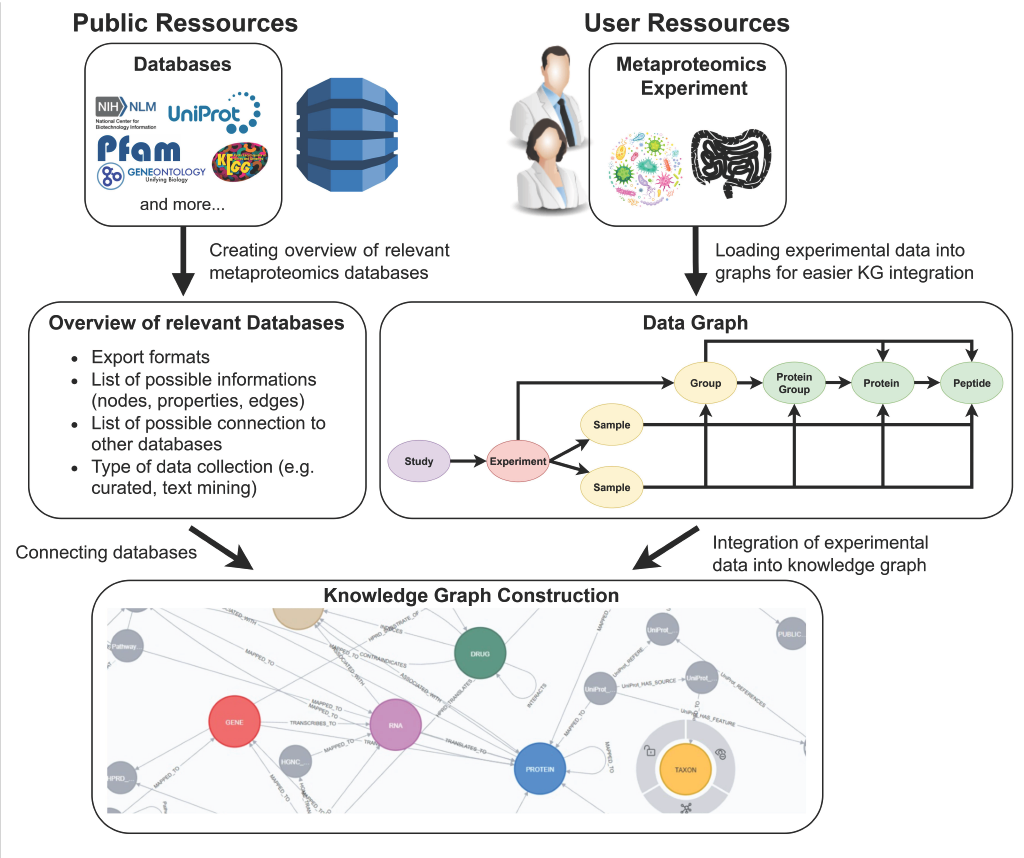
Brief summary of the main results and conclusion
The MetaProt-KG project established the foundation for a metaproteomics-centric knowledge graph to support microbiome multi-omics data integration and analysis. A result of the project was the systematic identification and cataloguing of relevant biological databases, with an initial focus on proteins, genes, taxonomy, and metabolites. Based on this assessment, a conceptual model for MetaProt-KG was developed and implemented as a Neo4j-based knowledge graph together with a prototype web application and database dump (distribution restricted for licensing reasons).
In parallel, a first collection of graph-based scripts and queries was created. These tools demonstrate how the knowledge graph can already support exploration of biological relationships and path-based analyses within its current scope. MetaProt-KG was presented at conferences and introduced in a hands-on metaproteomics workshop, showing its practical relevance and generating interest in further development.
The main conclusion is that the project demonstrated the feasibility and value of a metaproteomics-centered knowledge graph as a framework for microbiome research. At the same time, the work highlighted two important limitations: first, the current graph is still restricted by the availability and coverage of suitable databases across biological domains; second, licensing constraints from widely used resources such as KEGG and DrugBank currently prevent unrestricted public release of the full graph. These challenges do not diminish the project outcome, but rather define the next steps for sustainable expansion and dissemination.
Overall, MetaProt-KG provides a strong and expandable basis for future microbiome knowledge integration. The project delivered a working prototype, initial analytical functionalities, and community engagement activities that together position MetaProt-KG as a promising long-term resource. Future work will focus on extending the graph with additional biological domains, refining the analytical toolset, resolving licensing issues through compliant distribution strategies, and publishing the accompanying manuscript. In this way, MetaProt-KG can evolve from a metaproteomics-focused prototype into a broader community knowledge graph for microbiome research.
<< Back to all past Flex FundsProject Members
Prof. Dr. Robert Heyer
ORCID ID:
Leibniz-Institut für Analytische Wissenschaften - ISAS – e.V.
Prof. Dr. Dirk Benndorf
ORCID ID: 0000-0003-4021-8525
Anhalt University of Applied Sciences
Dr. Nico Jehmlich
ORCID ID: 0000-0002-5638-6868
Helmholtz Centre for Environmental Research GmbH – UFZ
Prof. Dr. Jana Seifert
ORCID ID: 0000-0002-7690-8539
University of Hohenheim
Keywords
Microbiom
Metaproteomics
Knowledge Graphs
Omics