Dear Reader,
With our 15th NFDI4Microbiota newsletter, we would like to share news about conferences, events, services and publications.
If you can think of other topics that we should cover, please let us know. We are happy to hear from you!
And now: Enjoy reading the newsletter!
Consortium News
NFDI4Microbiota is now on Matrix – Join our Community Chat

We are excited to announce that NFDI4Microbiota is now available on Matrix, offering a new and convenient way to connect with our community. Using the Element client, you can easily chat with us directly, ask questions, and share ideas.
We invite you to join our group, “NFDI4Microbiota Community,” where researchers, developers, and community members come together in an open and collaborative space. Whether you are looking for support or interested in networking, this channel provides a straightforward way to engage with us and others in the field.
Join the conversation here. We look forward to connecting with you!
Community engagement – calls, events and conferences
Call for contribution to shaping minimum standards for FAIR xenobiotics-microbiome biotransformation data submission
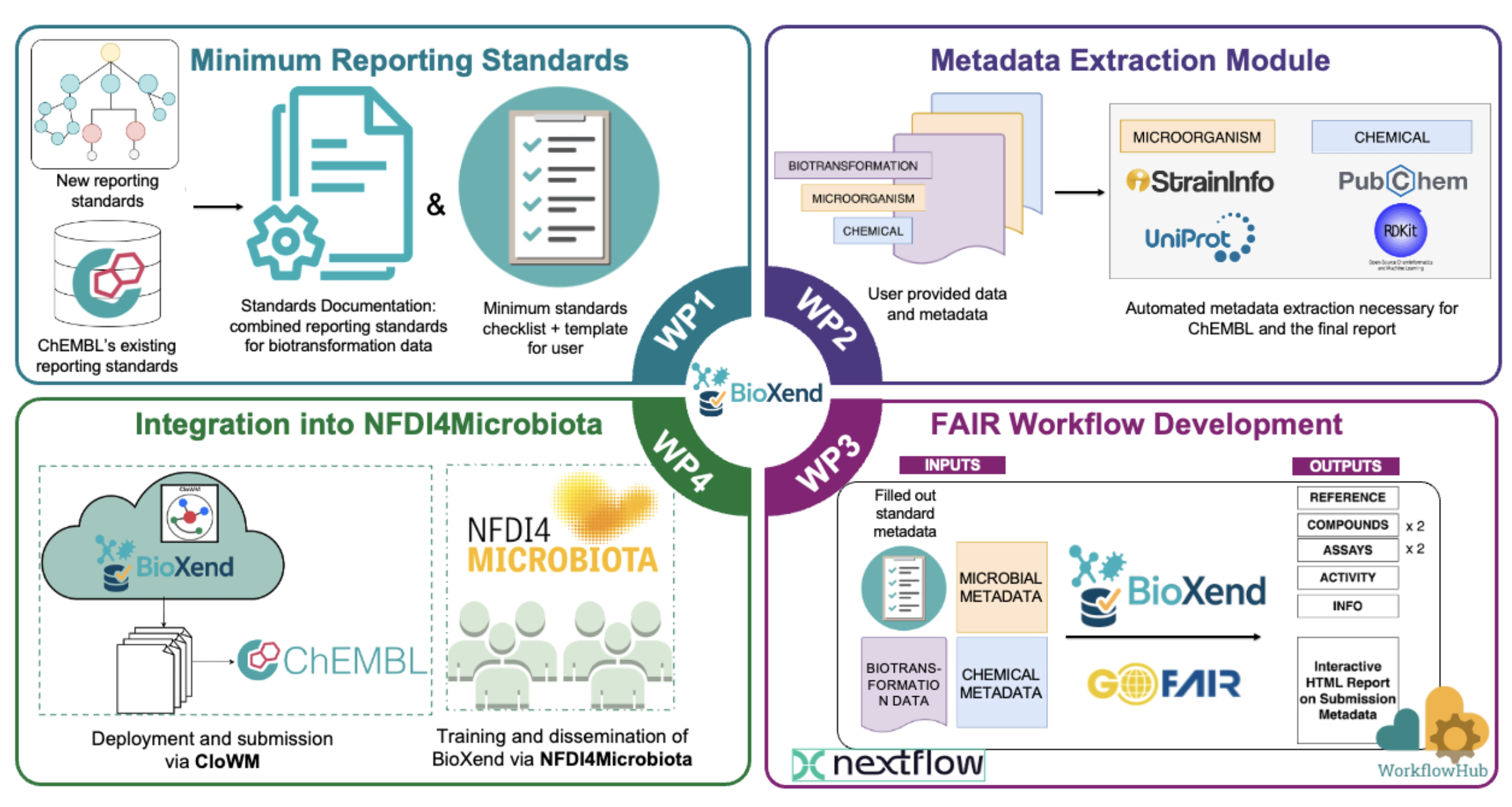
One of this year’s NFDI4Microbiota Flex Funds projects, “BioXend”, focuses on bridging the gap between microbiology and chemical biology. The project aims to develop a computational framework that enables FAIR and minimum standard compliant data submission to the ChEMBL database, specifically for xenobiotics–bacterial biotransformation data. To ensure these standards reflect the needs of the community, we invite you to contribute to this effort. You can get involved by:
- Participating in our open survey to share your expertise on data requirements
- Engaging on GitHub by submitting issues, opening pull requests, or starting discussions in the project repository
By contributing, you help streamline the submission process for microbiologists and lower the barriers to high-quality data sharing. We look forward to your input!
Submit your abstract for the NFDI4Microbiota Annual Conference

This year, NFDI4Microbiota will host its Annual Conference for the first time jointly with other consortia of the NFDI Biodata Interest Group: FAIRagro, DataPLANT, NFDI4Biodiversity, and NFDI4BIOIMAGE. The Boosting Biodata Bootcamp (B3 Conference) will take place from 15–17 September 2026 at RWTH Aachen University.
The aim of this joint event is to train the biological research community in research data management and data analysis, while providing inspiring insights into how these skills contribute to high-quality scientific research.
We invite life scientists — from laboratory-based to field researchers — as well as bioinformaticians, software developers, and local data stewards to share their FAIR tools and high-quality datasets, and to connect, exchange ideas, and discuss with experts in the field.
Participants will have the opportunity to learn how to use our services and make their data and tools FAIR. The conference program will include:
- Power Keynotes – Fresh impulses on bioinformatics, data management, and reproducible research
- Hands-on Training Sessions – Roll up your sleeves and work directly with tools, workflows & real datasets
- Lightning Talks – Fast-paced insights into cutting-edge methods, platforms, and real-world use cases
- Interactive Poster Arena – Exchange ideas, get feedback, spark collaborations
- Team-Up & Networking Sprints – Connect across disciplines
The call for abstracts is now open. Submit your abstract for a short talk or poster presentation by June 1, 2026.
Participants presenting a poster or talk at the conference are invited to apply for the FAIR Data Award during the abstract submission process. Submissions will be evaluated based on how well the underlying research data and analysis workflows adhere to good practices for data availability, documentation, and reusability.

We look forward to your participation!
Register for the 8th NFDI4Microbiota Community Workshop

On April 28, 13:00–14:30, we warmly invite you to the 8th NFDI4Microbiota Community Workshop, which will take place online. Under the motto “Looking back, looking forward: Practical tools to support your research”, we will reflect on past achievements and discuss how to further develop our services to better meet your needs.
As the first funding period ends in September 2026, this workshop also provides an opportunity to take stock of what has been achieved so far and to look ahead toward a second funding phase. Your input will be especially valuable in shaping the future direction of NFDI4Microbiota.
The workshop will focus on practical tools and solutions that directly support your research — from data management to analysis workflows.
During the workshop, we will provide a collaborative document where you can share your comments, challenges, and ideas. We are especially interested in the biggest problems we could help solve for you and which tools or services are currently missing.
Find more information and register here.
Join NFDI4Microbiota’s virtual Knowledge Base and Coding Sprint

Are you a microbiology researcher, research data management enthusiast or developer? We need your help! Join the NFDI4Microbiota Knowledge Base Sprint on April 22-23, 2026 online. The Knowledge Base offers extensive information about research data management, software development and much more. And you can help us to make this even better by contributing to the content or improving the code. Register here until April 15 to join the event online!
Collaborative hackathon at the International Virus Bioinformatics Meeting 2026
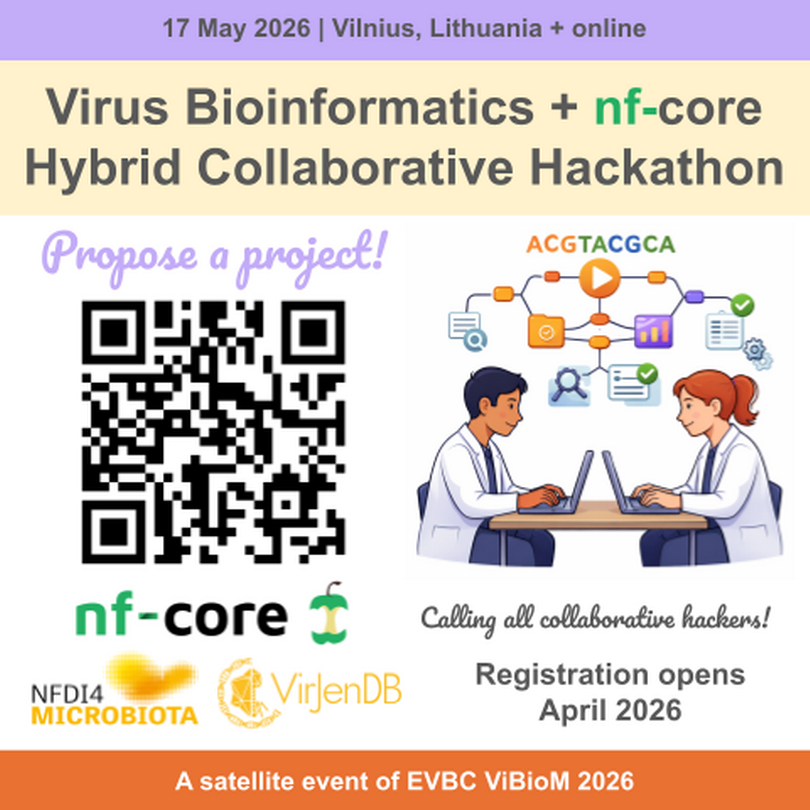

The VirJenDB team (NFDI4Microbiota) and the global nf-core community share a commitment to building open-source bioinformatics workflows (for example, in Nextflow) to enable reproducible research. We will organize a collaborative hackathon as a satellite event of the International Virus Bioinformatics Meeting 2026. You do NOT have to attend ViBioM to participate in the hackathon. We invite you to join us on May 17, 2026 in Vilnius, Lithuania, or online!
The intended audience are virus bioinformatics tool developers with an interest in reproducible workflows with skills in basic bioinformatics, e.g. bash, a programming language like python/R, and GitHub. We do welcome your hackathon project proposals! Please indicate your interest and/or propose a project in this survey. Registration for the hackathon will open in April 2026!
Recap on recent conferences

We were delighted to meet members of the microbial research community at our booth during the 2026 DGE Congress in March, held under the motto “Diet and the Microbiome – A Key to Good Health.” The event offered inspiring insights into nutrition research and its connection to the gut microbiome. We also took the opportunity to connect with scientists and other societies, including the Deutsche Gesellschaft für Mukosale Immunologie und Mikrobiom, while sharing our mission and services with a broader audience.
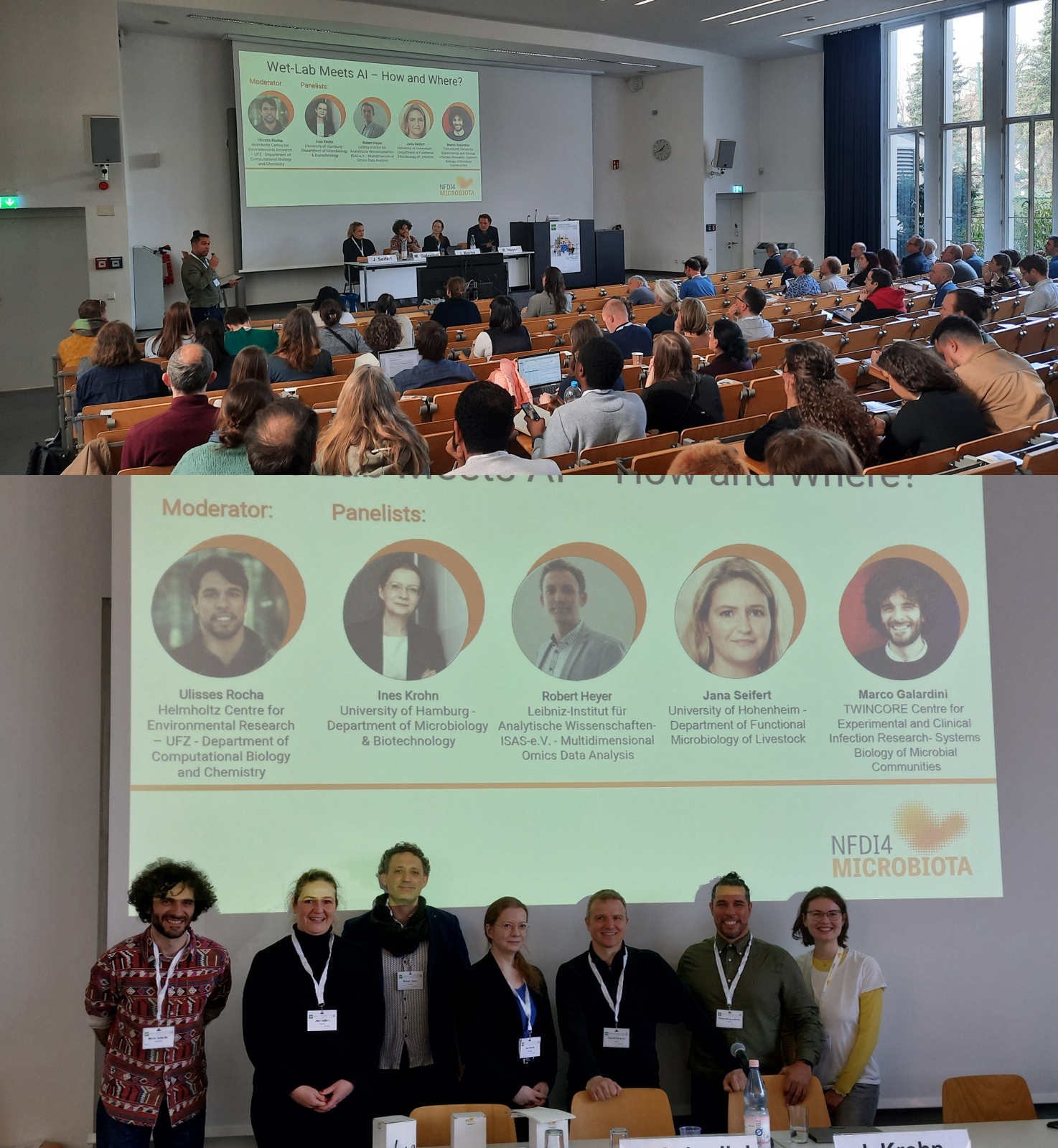

Once again, we participated in the Annual Conference of the Association for General and Applied Microbiology (VAAM) 2026, where we presented our work at a booth, during the poster session, and at our lunch symposium. A highlight of the symposium was the panel discussion titled “Wet-Lab Meets AI – How and Where?”, which brought together experts from experimental microbiology, applied microbiology, and computational biology to explore the current intersection of AI and wet-lab practices. The discussion addressed where AI-driven models are already influencing experimental planning, how multi-omics data and predictive modeling are transforming microbiological workflows, and where current approaches still face challenges when dealing with the complexity of microbial physiology. Our panelists—Ines Krohn (University of Hamburg), Robert Heyer (Leibniz Institute for Analytical Sciences – ISAS e.V.), Jana Seifert (University of Hohenheim), and Marco Galardini (TWINCORE Centre for Experimental and Clinical Infection Research)—engaged in a lively exchange moderated by NFDI4Microbiota partner Ulisses Rocha (Helmholtz Centre for Environmental Research).

Together, they discussed key questions such as the reliability of AI-based predictions in safety-relevant or applied contexts, the FAIRness and standardization of current microbiological datasets, and how researchers can balance mechanistic understanding with data-driven approaches.
Take a break and join our Coffee Talks

The Coffee Talk meetings provide an opportunity to learn more about NFDI4Microbiota’s services and mission from within and around the NFDI4Microbiota community. These bimonthly meetings take place online and are open to all who are interested in staying informed about current developments and topics related to data in microbiology research. Stay informed and find the schedule and details here.
Services
To support the microbial research community in data management and analysis, the NFDI4Microbiota consortium has established a range of dedicated services. Visit our persona webpage to explore which services are available for you as a wet-lab scientist, bioinformatician, data steward, or PI, or find an overview of all our services here.
Making Research Data More FAIR: Our Results from BioHackathon Germany 2025

Managing large-scale life science data in federated storage systems is challenging. While Research Object Crates (RO-Crate) offer a standardized way to package data with metadata, existing tools struggle with very large datasets and distributed environments. At BioHackathon Germany 2025, we addressed these challenges in our project “Enhancing FAIR (Meta-)Data Practices in Life Science by Improving RO-Crate Support in Federated Storage Systems.”
We focused on practical issues such as handling nested subcrates with complex references, merging independent RO-Crates while ensuring unique identifiers and resolving conflicts, and integrating checksum properties from the SPDX vocabulary to improve data integrity and validation. These discussions directly shaped our technical work throughout the week.
Over five days, a team of seven participants made significant progress, resulting in concrete improvements to the RO-Crate ecosystem and advancing more robust and FAIR-compliant data management practices. You can find more details about the results here.
Ask your questions to our Helpdesk!

The NFDI4Microbiota Helpdesk is the point of contact for the microbiology research community for all questions related to microbial (omics) data and associated metadata. We welcome questions from anyone - students, researchers, data stewards and more - working with this type of data, regardless of organism (bacteria, archaea, eukaryotic microorganisms, viruses), environment (e.g. soil, plants, host-associated and water) or data type (e.g. nucleic acid sequences, functional genomics, image data). Your question will be answered by one of the NFDI4Microbiota experts, including data stewards, researchers and software developers. You can contact us either by filling in our contact form or by email: helpdesk@nfdi4microbiota.de. We look forward to hearing from you!
Publications
NFDI4Microbiota’s FAIR data infrastructure enables the first truly planetary-scale tracking of microbial gene flow and antimicrobial resistance

We are excited to highlight how NFDI4Microbiota’s commitment to FAIR data has enabled a recent high-impact study published in the Journal Cell. By curating and integrating previously fragmented datasets, the NFDI4Microbiota-supported resources metaTraits (initiated as a NFDI4Microbiota Flex Fund) and Metalog, together with our partner EMBL’s SPIRE database of uniformly processed metagenomes, created an interoperable resource that enabled the first truly planetary-scale tracking of gene flow and antimicrobial resistance (AMR).
Spearheaded by Kim, Podlesny, and Schiller et al. from the Bork Group at EMBL Heidelberg, the study reveals fundamental structuring principles of the planetary microbiome. It demonstrates that a small group of ecologically highly adaptable “generalist” microbes can thrive broadly, connecting global microbiomes across diverse environments. Humans accelerate the dispersal of these microbes by creating new connections between environments that would otherwise remain separate. As they move between habitats, generalists can exchange genes with other microbes through horizontal gene transfer. This exchange can also spread AMR across environments, strongly highlighting the relevance of the One Health concept.
This milestone demonstrates how NFDI4Microbiota’s FAIR data efforts empower researchers to uncover complex connections across human, animal, and environmental health. DOI: https://doi.org/10.1016/j.cell.2025.12.051
VirJenDB: a FAIR (meta)data and bioinformatics platform for all viruses

One of NFDI4Microbiota services, VirJenDB, has been recently published in Nucleic Acids Research. The database VirJenDB offers a user-friendly, community-driven platform designed for versioned curation and ontology development. The database integrates virus sequences from 16 different sources, harmonizing nearly 200 metadata fields covering taxonomy, host, and sample information. So far, up to 85 metadata fields have undergone at least one round of curation, resulting in a resource that links 15.4 million viral sequences—88% from eukaryotic viruses and the remainder from prokaryotic viruses.
VirJenDB also provides curated subsets, including a novel collection of 0.91 million viral operational taxonomic units (vOTUs), while preserving original sequence diversity to support downstream analyses such as sequence variation studies.
The VirJenDB web portal offers intuitive access to both sequences and metadata via a search interface, filtering options, visualization tools, and an API for programmatic use. By enabling community contributions and supporting metadata standardization, VirJenDB aims to bridge phage and eukaryotic virus research communities, fostering interoperability, meta-analyses, and future tool integration. DOI: https://doi.org/10.1093/nar/gkaf1224